|
Hence, these results illuminate when exogenous factors and processes, such as chaperones or cotranslational folding, might be required for efficient protein folding.Synonyms for pervasive and other words similar to pervasive in our thesaurus. We also identify several properties and fold-types that are correlated with slow refolding on the minute time scale. Coli proteome is not intrinsically refoldable on physiological time scales, a cohort that is enriched with certain fold-types, domain organizations, and other biophysical features. Coli proteome expressed during log-phase growth, and among this group, we find that one-third of the E.(6−10) There are several possible reasons why a protein might not be able to refold to its native form. (4,5) These studies are typically interpreted through the ground truth of Anfinsen’s dogma, which states that proteins can intrinsically refold to their native states from unfolded forms because the native states represent global thermodynamic minima. Shangcheng Xu Cadmium (Cd) is a ubiquitous environmental and occupational.Many experiments of protein folding, conducted on purified, small, single-domain, soluble proteins, follow the proportion of protein molecules that are folded as a function of time, temperature, denaturant concentration, or sequence (1−3) and have yielded immense insight into the molecular determinants that underpin stable globular folds. Caldern-Aranda Abstract Lead (Pb) is a ubiquitous toxic metal. Obsessive-compulsive personality disorder is a pervasive pattern of preoccupation with orderliness, perfectionism, and mental and interpersonal control, at the expense of flexibility.
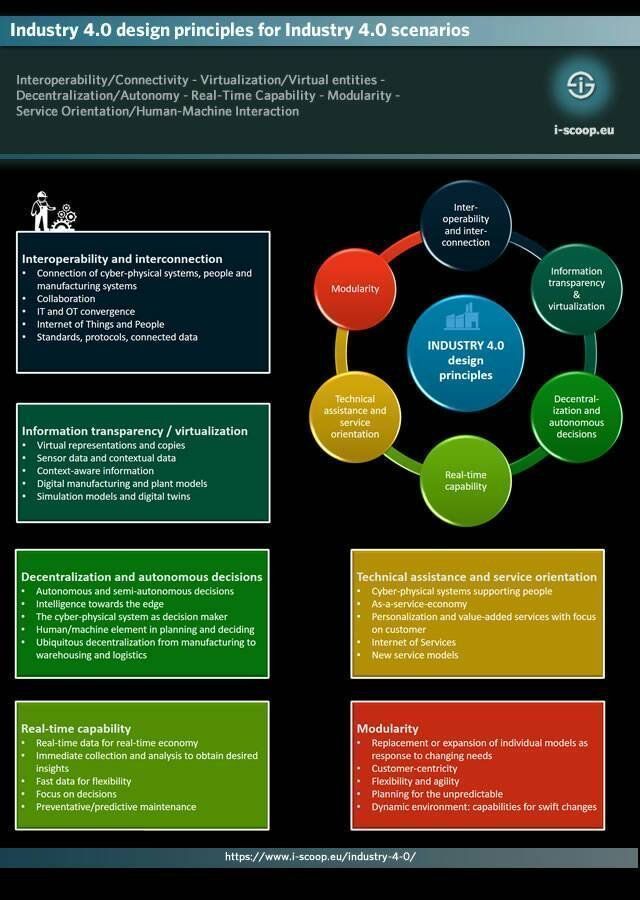
In this study, we introduce an experimental approach to probe protein refolding kinetics for whole proteomes. In any case, whether nonrefoldability is common for more complex “nonmodel” proteins is not generally known. (11,12) In other situations, it may be because the native state is challenging to access or is metastable relative to other folded (but non-native) conformations. 
Coli cells (type strain K-12) are grown in a defined media to the end of log phase (OD 0.8), resuspended in a lysis buffer, frozen by immersion into liquid nitrogen, and lysed by cryogenic pulverization (see Materials and Methods). This content is provided by iMedix and is subject to iMedix Terms.In our experimental design ( Figure 1), E. Autism affected children are also found to be mentally retarded. A low level of protease, active for a brief period of time (1 min), ensures that PK cleaves only at exposed or unstructured sites of target proteins, thereby encoding structural information on the protein’s conformation into cleavage sites. Following a period of time, the structures of the proteins in these complex mixtures are probed by subjecting them to pulse proteolysis with proteinase K (PK). (14) Clarified extracts are then divided so that one portion is retained in its native state, and a separate portion is first unfolded by addition of high concentrations of chemical denaturants (6 M guanidinium chloride (GdmCl)) and then refolded by lowering the denaturant concentration (by dilution or by dialysis). Ways to understand the meaning of the word is to know its synonyms.A reversibly refolding protein will have no “memory” of being unfolded, and hence following refolding, the protein will equilibrate to the same ensemble of conformations that it natively populated. Figure 1In business, something that is ubiquitous is widely adopted and can be found nearly. This method, called limited proteolysis mass spectrometry (LiP-MS), has been instrumental in exploring conformational change, (16) thermostability, (17) and allosteric binding (18) on the proteome-scale, and here, we have adapted it to probe protein refolding. By quantifying the relative abundance of HTPs arising from the refolded sample compared to those from the native sample, one can assess sites in a protein where the local conformation is different in the refolded form. Half-tryptic peptides (HTPs), in which one cut-site is deemed to arise from trypsin (which only cuts after Arg and Lys) and the other cut-site from PK (which can cut between any two residues), reveal a location where PK cleaved. Likewise, when refolded and native SNase are probed with LiP-MS, they generate a set of 147 distinct tryptic and half-tryptic peptides ( Figure 2B) that are all present in equal abundances within our cut-offs for significance (abundance ratio greater than 2-fold, P-value by Welch’s t test <0.01). SNase refolded by dilution out of 8 M urea has a CD spectrum that is superimposable with that of the native protein recombinantly expressed from E. To critically test these hypotheses, we first performed our LiP-MS method on two purified model proteins that are known refolders: Staphylococcal nuclease (SNase) and Ribonuclease H from Thermus thermophilus ( TtRNase H Figure S1A). We expect aggregated proteins to be more resistant to PK cleavage, while soluble misfolded proteins (with less compacted hydrophobic cores) are expected to be more susceptible to PK cleavage (cf. Super mario maker online course managerThese studies show that LiP-MS provides a consistent picture with CD for refolding proteins, although it does so with much greater structural resolution (providing independent quantifications at many distinct sites across the protein), and these studies show that complete refolding can be observed in a complex mixture. Coli lysate ( Figure 2F over 147 peptides). We repeated these studies on TtRNase H and found that its native and refolded forms generated overlapping CD spectra ( Figure 2D) and found no significant differences in their PK cleavage patterns, both in isolation ( Figure 2E over 176 peptides) and when spiked into E. Coli lysate ( Figure 2C), the conformations of native and refolded protein are again indistinguishable. Coli (green), and following unfolding in 8 M urea and 50-fold dilution (red). Circular Dichroism (CD) spectrum (A) of Staphylococcal nuclease (SNase) and (D) of Ribonuclease H from Thermus thermophilus ( TtRNase H), natively expressed from E. Refolding of Small Model Proteins. Battery blocsThe data suggest no significant difference in the structure of native SNase and the conformation produced when it is diluted out of urea. Inset shows large number of points clustered near the origin. Red regions designate significance (effect-size >2, P-value <0.01). Effect sizes reported as ratio of averages, and P-values are based on Welch’s t test. (B) Volcano plot comparing peptide abundances from native and refolded SNase ( n = 3). We reasoned that ribosomal proteins and RNA, which are highly abundant and charged, might potentially be the most prone to aggregate and seed the aggregation of other proteins. Coli lysate.Nevertheless, pelleting assays showed that in these complex mixtures, still ∼11.5% of the total protein content is precipitated after refolding reactions ( Figure S2B). (F) As in part C, except for TtRnaseH spiked into E. (E) As in part B, except for the purified protein, TtRnaseH. Coli lysate, providing a complex background. 
0 Comments
Leave a Reply. |
AuthorHeather ArchivesCategories |
 RSS Feed
RSS Feed
